 |
Ziyang Tan received the online comment from AIP “Reducing background signals in SERS detection and imaging”
In recent years, nanoprobes called surface-enhanced Raman scattering (SERS) tags have been developed for detection of biological matter and imaging applications. These nanoscale probes combine metallic nanoparticles and specific organic Raman reporter molecules to sense target molecules with laser Raman spectroscopy or SERS.
One challenge for these devices, however, is the high background signals from cell culture containers’ or biological tissues’ autofluorescence, which can mask signals from the SERS tags. Tan et al. used mathematical methods to mitigate the influence of these unwanted background signals. Their results suggest that, as long as the intensity of specific Raman bands can overcome noise of the whole spectrum, biological applications of SERS tags should not have problems with unwanted background signals.
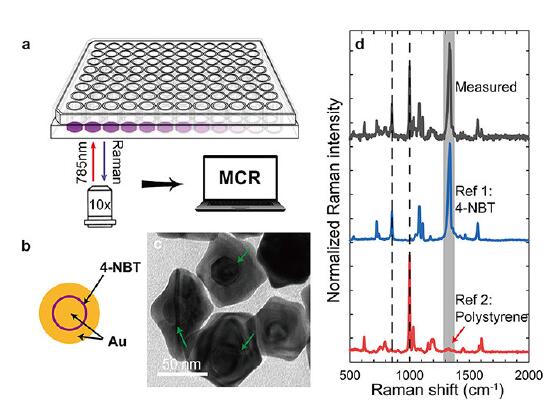 To test their theory, the researchers synthesized gold gap-enhanced Raman tags (GERTs), a new type of SERS tag with a subnanometer-sized interior gap between a metallic core and shell. The tags were deposited in each well of a 96-well plate, with laser excitation and Raman collection occurring from the bottom side. For autofluorescence measurements, human lung cancer cells were adhered to quartz substrates and then incubated with GERTs before performing multiplex Raman imaging.
To test their theory, the researchers synthesized gold gap-enhanced Raman tags (GERTs), a new type of SERS tag with a subnanometer-sized interior gap between a metallic core and shell. The tags were deposited in each well of a 96-well plate, with laser excitation and Raman collection occurring from the bottom side. For autofluorescence measurements, human lung cancer cells were adhered to quartz substrates and then incubated with GERTs before performing multiplex Raman imaging.
For both SERS detection and imaging, the team compared the conventional peak integral method and two multivariate curve resolution (MCR) methods: negative matrix factorization and classical least squares. The results demonstrate that MCR methods lower the limit of detection by one order of magnitude during SERS detection at high Raman background compared to the peak integral method. For SERS imaging with autofluorescence background, the new methods significantly eliminated both false-negative and false-positive results.